Click here to see a list of programs developed by the partners of the platform.
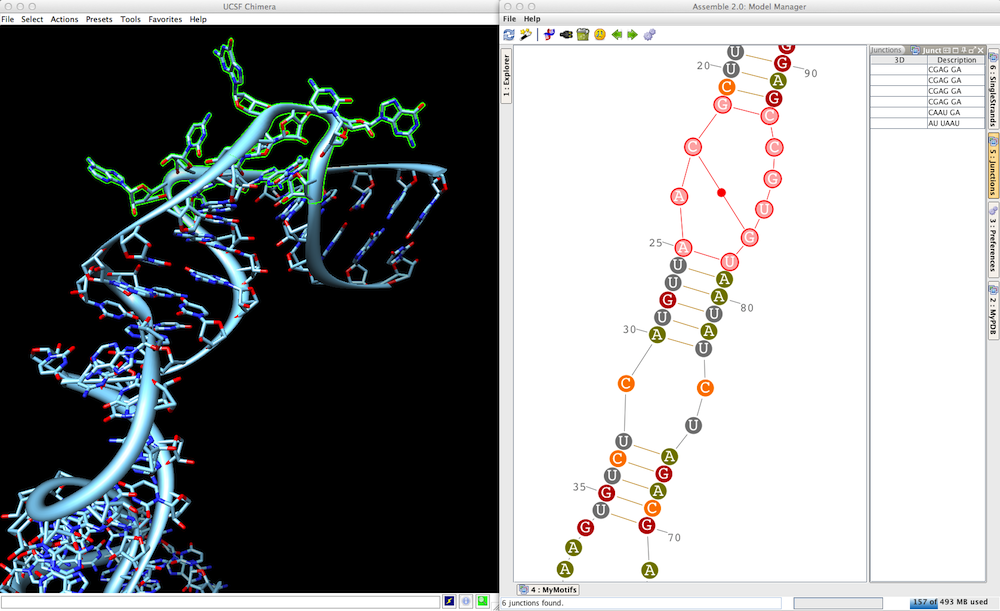 | Assemble 2: Study and construct RNA 3D architectures interactively. | DOI: 10.1093/bioinformatics/btq321 |
 | MACSIMS: Multiple Alignment of Complete Sequences Information Management System. | DOI: 10.1186/1471-2105-7-318 |
 | Rapid detection of orthology and inparalogy relations. | DOI: 10.1093/nar/gky1068 |
 | Quality control systems for ChIP-seq and enrichment-related datasets. | DOI: 10.1093/nar/gkt829 |
 | Comprehensive workflow for annotating and ranking SNVs and indels from next generation sequencing data. | DOI: 10.7717/peerj.796 |
 | MS/MS spectra viewer/extractor designed to extract “high quality” spectra from peaklist files. | DOI: 10.1002/pmic.201300415 |
 | Data integration framework and software suite for mass spectrometry based proteomics | http://www.profiproteomics.fr/proline/ |
 | AnnotSV: An integrated tool for Structural Variations annotation. | DOI: 10.1093/bioinformatics/bty304 |
 | Mychem is a chemoinformatics extension for MySQL and MariaDB. | https://mychem.github.io/ |
 | Highlight protein divergences between species | DOI: 10.1093/gbe/evaa248 |
 | A prediction tool to reveal disease-relevant deleterious missense variants. | DOI: 10.1371/journal.pone.0236962 |
